(The Human Genome Project) opened the door to a vast labyrinth of new questions (…) the complexity of biology has seemed to grow by orders of magnitude.
Hayden (2010)
If the Human Genome Project gave us a book, scientists are now learning how to read it (...) and biologists are beginning to face up to the uncomfortable truth that they have only been looking at the nouns (...) now we are reading the spaces in between—verbs, adverbs, adjectives, pronouns and the rest, and they are complicated indeed.
Roger Highfield1
By giving us a catalogue of the most basic elements of human biology – genes – the completion of the Human Genome Project (HGP) marked the beginning of a new era for medicine. Armed with this knowledge, scientists would be able to correlate the manifestation of diseases and genetic mutations. In turn, the pharmaceutical industry would develop therapies by selectively targeting mutations.
Based on this premise, many scientists and observers predicted a swift therapeutical revolution. For example, Randy Scott of Incyte Genomics stated that “in 10 years, we will understand the molecular basis for most human diseases” (Palmer 2013), while President Clinton claimed that this knowledge would “revolutionise the diagnosis, prevention and treatment of most, if not all, human diseases”.2
In the 15 years since the HGP’s completion, medicine has experienced a significant transformation at the hand of genetics. Examples of this transformation include the rapid development of basic research fields such as genetic epidemiology and pharmacogenomics, the incorporation of genetics to the core curricula of leading medical programmes, and the emergence of genetic diagnostics and personalised medicine. Additionally, being able to probe for the interaction of compounds and specific mutations has streamlined therapeutical innovation by reshaping the way in which drug targets and clinical trial subjects are selected.
Nevertheless, impatience and criticism have arisen based on the idea that this ‘genetic revolution’ has not yet materially improved patient outcomes, failing to have met the more optimistic expectations of 15 years ago (Varmus 2010). This criticism highlights that few game-changing genetic therapies have reached the market (Mardis 2011), that disease mortality continues to be largely driven by the same causes of two decades ago, and that the molecular basis for most important diseases remains to be fully elucidated (Wade 2010, Palmer 2013). Some have questioned the worthiness of the ‘genetic endeavour’ (Evans et al. 2011), even suggesting that a ‘genomic bubble’ may have “temporarily bogged down the drug industry with information overload” (Pollack 2010).
In recent work (Hermosilla and Lemus 2017) we seek to contribute to this debate by analysing the extent to which basic genetic science has fuelled (or ‘translated into’) early-stage drug innovation. Our evidence is consistent with the idea that the alleged ‘slower-than-expected’ progress has been partly caused by the large amount of complexity in human biology, which, as suggested by the opening quotes, was unexpected prior to the Project’s completion and has been progressively revealed since then.
Biological complexity
Over the last ten years, scientific research has produced a number of results that illustrate the astounding complexity of human biology. For example, we now know that genetic mutations rarely map one-to-one into diseases (Bauer-Mehren et al. 2011), that non-protein-coding genes may regulate protein-coding ones (Li et al. 2016), and that common genetic mutations explain a relatively small percent of predicted genetic variance between individuals (Manolio et al. 2009). The recent ‘omnigenic’ hypothesis (Boyle et al. 2017) further complicates matters by suggesting that seemingly unrelated ‘peripheral’ genes (located outside core pathways, and which cannot be easily categorised based on known biology) may drive disease through cellular networks.
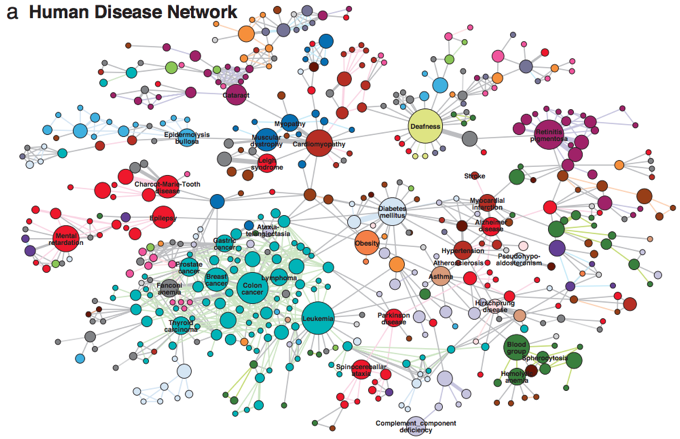
The Human Disease Network (Goh et al. 2007) highlights one manifestation of this complexity that is important to understand therapeutic translation. In this network, diseases are nodes, while common mutations between two diseases are edges (or connectors). Figure 1 describes the network graphically.3
Figure 1 A Human Disease Network
Source: Figure 2a in Goh et al. (2007).
Some diseases in this network (like colon cancer) are connected to many others. Compared to other diseases that are scantily connected (like myopathy), translating basic genetic discoveries into therapeutics that target the former type of disease may be more difficult. There are at least two reasons:
- One, the large number of associated mutations may make ‘silver bullet’ therapies harder to craft.
- And two, therapies targeting highly-connected diseases grapples with a higher risk of interfering with related biological processes, and so a higher risk of producing adverse side effects.
We implemented an updated representation of the Human Disease Network and used it to construct measures of complexity based on the above notion. Consistent with Figure 1, the resulting variables reveal substantial heterogeneity in the biological complexity of different diseases.4 Focusing on the ten years that followed the Project’s completion, we exploited this longitudinal variation by estimating its dependency on a disease-level measure accumulated genetic-epidemiological knowledge.
Therapeutic translation in the wake of genome
Our empirical analysis amounts to deciphering whether diseases’ measured complexity mediates the rate of translation of new genetic epidemiological findings into early-stage drug innovation.
We implemented this analysis by focusing on the translation of a leading type of genetic epidemiological research, namely, the genome-wide association studies (GWASs). These studies search for genetic mutations underlying the manifestation of diseases, validating associations only if stringent statistical significance standards are met.5 Despite having a number of limitations (e.g. Cao and Moult 2013), the GWA approach has “played a major role in unravelling the genetic bases of complex diseases” (So et al. 2017) and “opened doors to potential treatments by revealing the unexpected involvement of certain functional and mechanistic pathways in a variety of disease processes” (Manolio 2010).
We complemented GWAS publication data with a large pharmaceutical pipelines dataset, which tracks the number of new therapies that entered the pre-clinical development stage (2003-2012) at the targeted-disease level. We then estimated the rate of translation of accumulated GWAS knowledge using standard methods for the measurement of R&D productivity (Blundell et al. 2002).
For less complex diseases, we found a strong and positive association between cumulative GWAS knowledge and the amount of innovation. This association weakens as complexity increases in a roughly monotonic fashion, tracing a negative ‘complexity/translation rate’ gradient. Furthermore, the association becomes statistically insignificant among diseases associated with the highest measured complexity. We take these findings as evidence for the idea that biological complexity is in part responsible for the slower-than-expected unfolding of the therapeutic revolution set in motion by the Human Genome Project.
Relevance
Despite a rich stream of research investigating the extent to which basic science fuels and shapes pharmaceutical innovation, biological complexity has been invariably treated as ‘unobserved heterogeneity’ by systematic analyses. This omission stands out as particularly relevant in the new ‘genomic era’, where abundant biological data may inform development decisions.
Furthermore, the documented ‘complexity/translation rate’ gradient suggests that complexity may be an important conditioning factor in order to evaluate pharmaceutical R&D productivity. For scientific funding agencies, it may suggest that applied research focusing on ‘less complex’ diseases may be more likely to have a therapeutic impact in the short term.
References
Bauer-Mehren, A, M Bundschus, M Rautschka, M A Mayer, F Sanz and L I Furlong (2011), “Gene-disease network analysis reveals functional modules in mendelian, complex and environmental diseases”, PloS one 6(6): e20284.
Blundell, R, R Griffith, and F Windmeijer (2002), “Individual effects and dynamics in count data models”, Journal of Econometrics 108(1): 113–131.
Boyle, E A, Y I Li, and J K Pritchard (2017), “An expanded view of complex traits: From polygenic to omnigenic”, Cell 169(7): 1177–1186.
Cao, C and J Moult (2014), “GWAS and drug targets”, BMC genomics 15(4): S5.
Goh, K I, M E Cusick, D Valle, B Childs, M Vidal & A L Barabási (2007), “The Human Disease Network”, Proceedings of the National Academy of Sciences 104(21): 8685-8690.
Hayden, E (2010), “Human genome at ten: life is complicated”, Nature News 464(7289): 664-667.
Evans, J P, E M Meslin, T M Marteau, and T Caulfield (2011), “Deflating the genomic bubble”, Science 331(6019): 861–862.
Hermosilla, M I and J A Lemus (2017), “Therapeutic Translation in the Wake of the Genome”, NBER Working Paper No. 23989.
Li, Y I, B van de Geijn, A Raj, D A Knowles, A A Petti, D Golan, Y Gilad, and J K Pritchard (2016), “RNA splicing is a primary link between genetic variation and disease”, Science 352(6285): 600–604.
Manolio, T A (2010), “Genomewide association studies and assessment of the risk of disease”, New England Journal of Medicine 363(2): 166-176.
Manolio, T A (2013), “Bringing genome-wide association findings into clinical use”, Nature Reviews Genetics 14(8): 549.
Manolio, T A, F S Collins, N J Cox, D B Goldstein, L A Hindorff, D J Hunter, M I McCarthy, E M Ramos, L R Cardon, A Chakravarti, et al. (2009), “Finding the missing heritability of complex diseases”, Nature 461 (7265): 747–753.
Mardis, E R (2011), “A decade’s perspective on DNA sequencing technology”, Nature 470(7333): 198.
Palmer, B (2013), “Where are all the miracle drugs?” Slate, 30 September.
Pollack, A (2010), “Awaiting the genome payoff”, The New York Times, 14 June.
So, H C, C K L Chau, W T Chiu, K S Ho, C P Lo, S H Y Yim, and P C Sham (2017), “Analysis of genome-wide association data highlights candidates for drug repositioning in psychiatry”, Nature Neuroscience 20(10): 1342.
Wade, N (2010), “A decade later, genetic map yields few new cures”, The New York Times, 12 June.
Endnotes
[1] Highfield is a former Editor of New Scientist; see “Life just got a lot more complicated,” The Telegraph, 19 June 2007.
[2] See “President Clinton, British Prime Minister Tony Blair Deliver Remarks on Human Genome Milestone,” CNN, 26 June 2000.
[3] This figure is reproduced from Goh et al. (2007).
[4] Our implementation uses the over half-a-million gene-disease associations that are available from DisGeNet, a publicly available and systematic repository of genetic epidemiological results. The resulting representation is much more dense (and its graph much less informative) than that of Goh et al. (2007).
[5] GWASs started to be published in 2005. They comprise a tiny fraction of the gene-disease association results available from DisGeNet.